SegTraQ#
SegTraQ (Segmentation and Transcript Assignment Quality Control) is a Python toolkit for quantitative and visual quality control of segmentation and transcript assignment in spatial omics data.
⚠️ Note: SegTraQ is under active development. Features, interfaces, and functionality may change in upcoming releases.
Contents:
Tutorials:
- Data Preparation
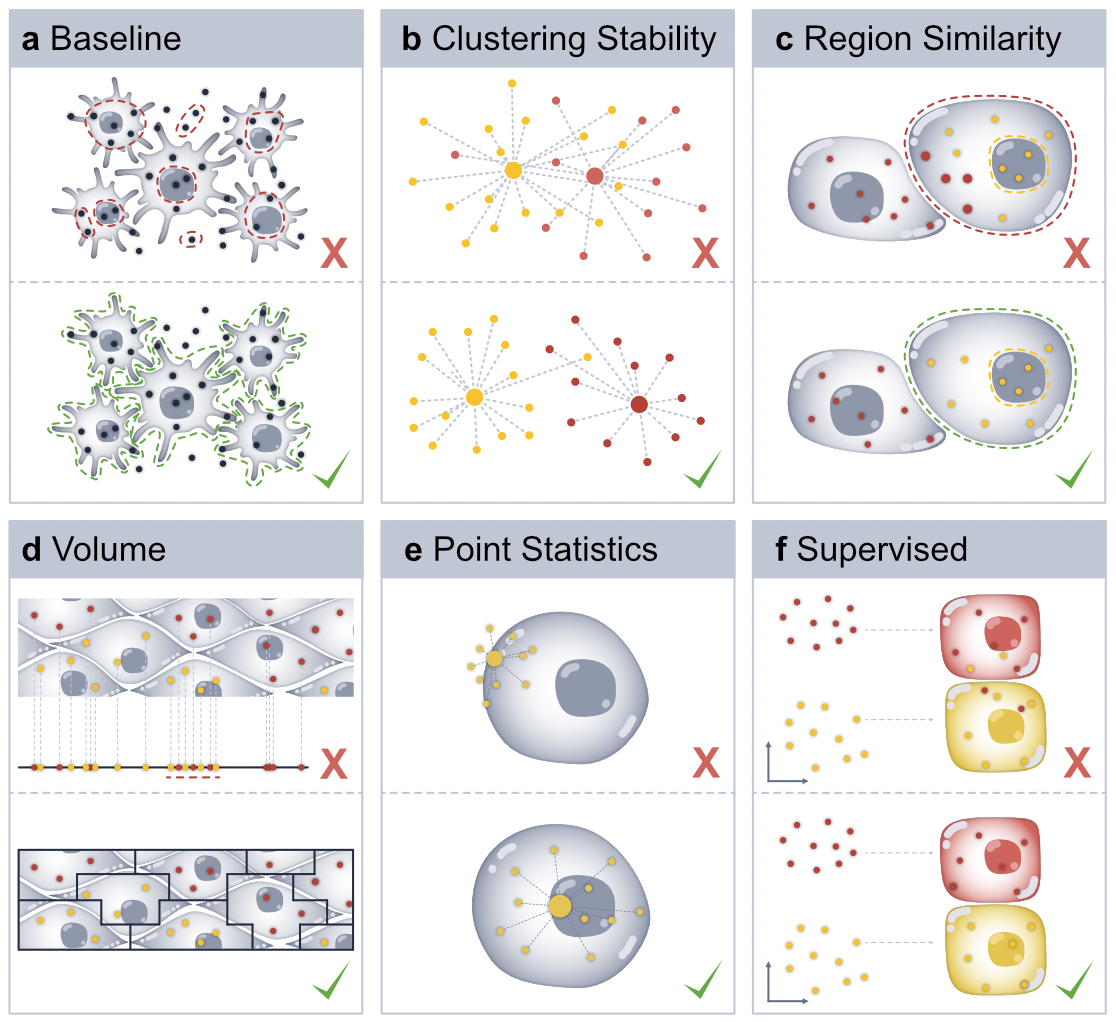
- Module: Baseline Metrics
- Module: Clustering Stability
- Module: Region Similarity
- Module: Volume (3D)
- Module: Supervised Metrics
- Module: Point Statistics
- Module: Plotting
- Plots for individual samples
- Comparing between segmentation methods
- Technology Focus: 10x Genomics Xenium (Simplified)
- Technology Focus: 10x Genomics Xenium

